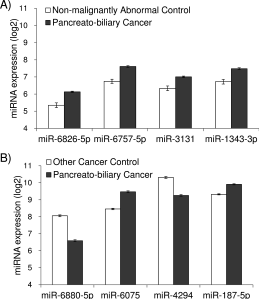
A recent paper by Kojima, M. et al. has found a signature of miRNAs to identify patients with pancreato-biliary cancers who could benefit from surgical intervention.
Namely, a combination of eight miRNAs (miR-6075, miR-4294, miR-6880-5p, miR-6799-5p, miR-125a-3p, miR-4530, miR-6836-3p, and miR-4476) achieved a sensitivity, specificity, accuracy and AUC of 80.3%, 97.6%, 91.6% and 0.953, respectively.
In contrast, CA19-9 and CEA gave sensitivities of 65.6% and 40.0%, specificities of 92.9% and 88.6%, and accuracies of 82.1% and 71.8%, respectively, in the same test cohort. This diagnostic index identified 18/21 operable pancreatic cancers and 38/48 operable biliary-tract cancers in the entire cohort.
Finding of this eight miRNAs was possible by using Toray’s 3D profiling technology. This finding is especially important, as it is difficult to detect pancreatic cancer or biliary-tract cancer at an early stage using current diagnostic technology.
microRNAs are stably present in peripheral blood, and are therefore a good candidate for finding prognostic or diagnostic biomarkers.
Studying only the miRNAs available in the literature may limit the novelty of the biomarkers found, so for a wide variety of diseases, a profiling is mandatory in order to find really specific circulating biomarkers.
Toray’s technology is available in Europe as fee-for-services via tebu-bio Laboratories (see Press Release). By exploring the full miRnome (composed by 2,000 miRNAs) with Toray’s 3D-Gene® technology, one might identify slight miRNA expression level changes (including low abundance miRNAs) in blood samples, biopsies FFPE specimens…
Interested in miRNA profiling in cancer research? Leave a message below or browse tebu-bio’s 3D-Gene® miRNA profiling platform web page!