Following on in our series on microRNAs (miRNA – Keys to open cell biology’s secret chamber) , in this post, we’ll be taking a look at Antagomirs (which are also called anti-miRs or blockmirs). These chemically engineered oligonucleotides prevent other molecules from binding to a desired site on an mRNA molecule. They’re used to silence endogenous microRNA.
Antagomirs & Antagomir database
Owing to several pitfalls inherent to antagomirs (toxicity, tissue distribution and pharmacokinetics issues), it will be difficult to predict their effectiveness until they’re empirically tested in vitro. There are few databases which can be helpful when designing antagomirs, one of the most popular is AntagomirBase .
Luckily, as they are becoming very popular tools, the literature can be of great help to find an appropriate antagomir.
As for miRNA mimics, chemically modified RNAs (capped RNA molecules) can be a good starting choice as antagomirs in very short term experiments. (LNA vs. 2’-O-Me oligonucleotides)
Plasmids coding for antagomirs
This is the most classical approach to keep inhibition of the miRNA (miRNA arrest) for long term experiments. With an appropriate selection and screening method, you can obtain a polyclonal population of cells expressing low, medium and high antagomir levels (for comparative studies or other applications). A broad selection of vectors is available, for you to select the most appropriate for your research.
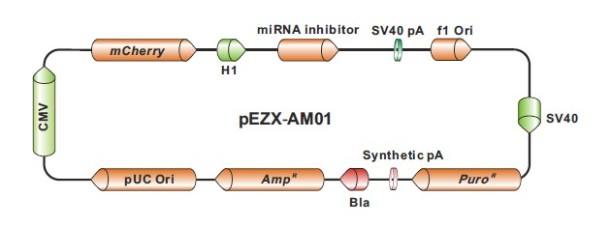
Lentiviral vectors
Again, some primary cultures or established cell lines can be difficult to transfect, but this difficulty can be overcome by using lentiviral tools. However, the use of lentiviral tools will lead to homogeneous expression of the antagomir in cells (“monoclonal-like” cell lines).
As always, don’t hesitate to ask your questions…
Just leave any comments or questions below, I’ll be pleased to answer!