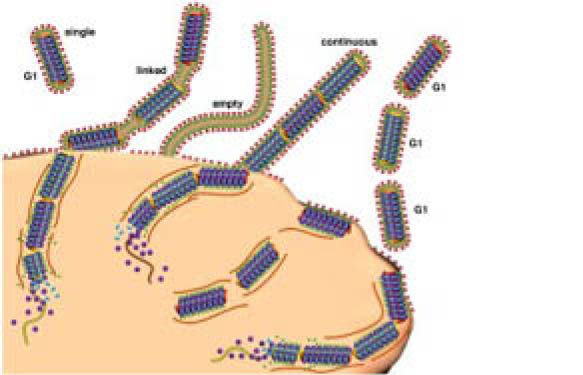
In a previous post on whether samples should be pooled or not for proteomic profiling, we discussed this approach, which can be quite cost-wise, while still allowing to see the main biomarkers differentiating one physiological condition from another (e.g. disease vs healthy control).
In real life, however, this discrimination between physiological conditions may be difficult to define. Let’s take, for example, a study aimed at studying the differential immune response to an infection, and how this can be used to design more efficient therapies in different population subgroups.
There are several criteria that must be taken into account:
Pathogens can be quite different. Viral serotypes, bacterial strains, parasite species and sub-species… Depending on the serotype / strain / species / sub-species, the clinical symptoms will vary… and so will the relevant biomarkers.
It’s therefore important to make sure that we have thoroughly characterised the pathogen involved in the infection, and what can be generically indicated as “flu” represents in fact a homogeneous infection by a given serotype.
If we are not sure about this homogeneity, then pooling is not recommended. Pooling should only be performed on a homogeneous group of samples.
Criteria #2 – what patient?
How can we define homogeneity in a group of patients? Who is healthy and who has a disease?
If we take a simple approach, disease vs healthy will be defined by a given condition (usually the one we want to study). However, human beings are not (normally) ill for one single cause. There are other health-associated status that can complicate the search for biomarkers.
Let’s take a patient with, say, a viral infection. This patient, apart from this viral infection, may also have other health conditions such as obesity, diabetes, cardiovascular diseases, etc. This may complicate the analysis for biomarkers.
In general, if we want to get rid of this “background” disease, we will ignore them and pool samples depending on the infection, and not the health status of each individual.
However, if we want to study, precisely, how background health status affects the response to a given infection or therapy, we should them group individuals with similar background health status…or not pool at all.
Pooling will allow us to identify generic biomarkers for a given condition. Not pooling will allow us to study the differences between the individuals, and maybe find new subgroups (e.g. responders vs non-responders).
It depends on whether you want to concentrate on studying the trees or see the whole forest losing the details. Each project will require a different approach, and at the Biomarkers team at tebu-bio, we will be glad to support you in this process that can affect the outcome of your research dramatically.
This is a topic of continuous debate, so stay tuned! We would love to hear your feedback!